
What is discovr?
The discovr package will contain tutorials associated with my textbook
Discovering Statistics using R and RStudio, due out in early 2021. It will include all datasets, but most important it will contain a series of interactive tutorials that teach
alongside the chapters of the book. The tutorials are written using a package called
learnr. Once a tutorial is running it’s a bit like reading excerpts of the book but with places where you can practice the R code that you have just been taught. The discovr package is free (as are all things
-related) and offered to support tutors and students using my textbook.
What are R and RStudio?
If you’re using a textbook about then you probably already know what it is. If not, R is a free software environment for statistical analysis and graphics. RStudio is a user interface through which to use . RStudio adds functionality that make working with easier, more efficient, and generally more pleasant than working in alone.
You can get started with R and RStudio by completing this tutorial (includes videos):
Contents of discovr
The tutorials are named to correspond (roughly) to the relevant chapter of the book. For example, discovr_04 would be a good tutorial to run alongside teaching related to chapter 4, and so on. Some longer chapters have several tutorials that break the content into more manageable chunks. Given the current global situation and the fact that lots of instructors are needing to teach remotely I’m making what I have available before the book is finished and will update as and when new tutorials are written.
- discovr_01: Key concepts in (functions and objects, packages and functions, style, data types, tidyverse, tibbles)
- discovr_02: Summarizing data (frequency distributions, grouped frequency distributions, relative frequencies, histograms, mean, median, variance, standard deviation, interquartile range)
- discovr_03: Confidence intervals: interactive app demonstrating what a confidence interval is, computing normal and bootstrap confidence intervals using , adding confidence intervals to data summaries.
- discovr_05: Visualizing data. The ggplot2 package, boxplots, plotting means, violin plots, scatterplots, grouping by colour, grouping using facets, adjusting scales, adjusting positions."
- discovr_06: The beast of bias. Restructuring data from messy to tidy format (and back). Spotting outliers using histograms and boxplots. Calculating z-scores (standardizing scores). Writing your own function. Using z-scores to detect outliers. Q-Q plots. Calculating skewness, kurtosis and the number of valid cases. Grouping summary statistics by multiple categorical/grouping variables.
- discovr_07: Associations. Plotting data with GGally. Pearson’s r, Spearman’s Rho, Kendall’s tau, robust correlations.
- discovr_08: The general linear model (GLM). Visualizing the data, fitting GLMs with one and two predictors. Viewing model parameters with broom, model parameters, standard errors, confidence intervals, fit statistics, significance.
- discovr_09: Categorical predictors with two categories (comparing two means). Comparing two independent means, comparing two related means, effect sizes.
- discovr_10: Moderation and mediation. Centring variables (grand mean centring), specifying interaction terms, moderation analysis, simple slopes analysis, Johnson-Neyman intervals, mediation with one predictor, direct and indirect effects, mediation using
lavaan. - discovr_11: Comparing several means. Essentially ‘One-way independent ANOVA’ but taught using a general linear model framework. Covers setting contrasts (dummy coding, contrast coding, and linear and quadratic trends), the F-statistic and Welch’s robust F, robust parameter estimation, heteroscedasticity-consistent tests of parameters, robust tests of means based on trimmed data, post hoc tests, Bayes factors.
- discovr_12: Comparing means adjusted for other variables. Essentially ‘Analysis of Covariance (ANCOVA)’ designs but taught using a general linear model framework. Covers setting contrasts, Type III sums of squares, the F-statistic, robust parameter estimation, heteroscedasticity-consistent tests of parameters, robust tests of adjusted means, post hoc tests, Bayes factors.
- discovr_13: Factorial designs. Fitting models for two-way factorial designs (independent measures) using both
lm()and theafexpackage. This tutorial builds on previous ones to show how models can be fit with two categorical predictors to look at the interaction between them. We look at fitting the models, setting contrasts for the two categorical predictors, obtaining estimated marginal means, interaction plots, simple effects analysis, diagnostic plots, partial eta-squared and partial omega-squared, robust models and Bayes factors. - discovr_14: Repeated measures designs. Fitting models for one- and two-way repeated measures designs using the
afexpackage. This tutorial builds on previous ones to show how models can be fit with one or two categorical predictors when these variables have been manipulated within the same entities. We look at fitting the models, setting contrasts for the categorical predictors, obtaining estimated marginal means, interaction plots, simple effects analysis, diagnostic plots, robust models and Bayes factors. - discovr_15: Mixed designs. Fitting models for mixed designs using the
afexpackage. This tutorial builds on previous ones to show how models can be fit with one or two categorical predictors when at least one of these variables has been manipulated within the same entities and at least one other has been manipulated using different entities. We look at fitting the models, setting contrasts for the categorical predictors, obtaining estimated marginal means, interaction plots, simple effects analysis, diagnostic plots, robust models and Bayes factors. - discovr_18: Exploratory Factor Analysis (EFA). Applying factor analysis using the
psychpackage. This tutorial uses a fictitious questionnaire (the Anxiety Scale, RAQ) with 23 items to show how EFA can be used to identify clusters of items that may, or may not, represent constructs associated with anxiety about using . We look at inspecting the correlation matrix, obtaining the Bartlett test and KMO statistics, using parallel analysis to determine the number of factors to extract, extracting factors, rotating the solution nd interpretation of the factors. We also learn to obtain Cronbach’s alpha on each of the subscales.
Installing discovr
To use discovr you first need to install
and RStudio. To learn how to do this and to get oriented with
and RStudio complete my interactive tutorial,
getting started with R and RStudio.
You can get the development version of the package from github.com/profandyfield/discovr.
Running a tutorial
In RStudio Version 1.3 onwards there is a tutorial pane. Having executed
library(discovr)
A list of tutorials appears in this pane. Scroll through them and click on the  button to run the tutorial:
button to run the tutorial:
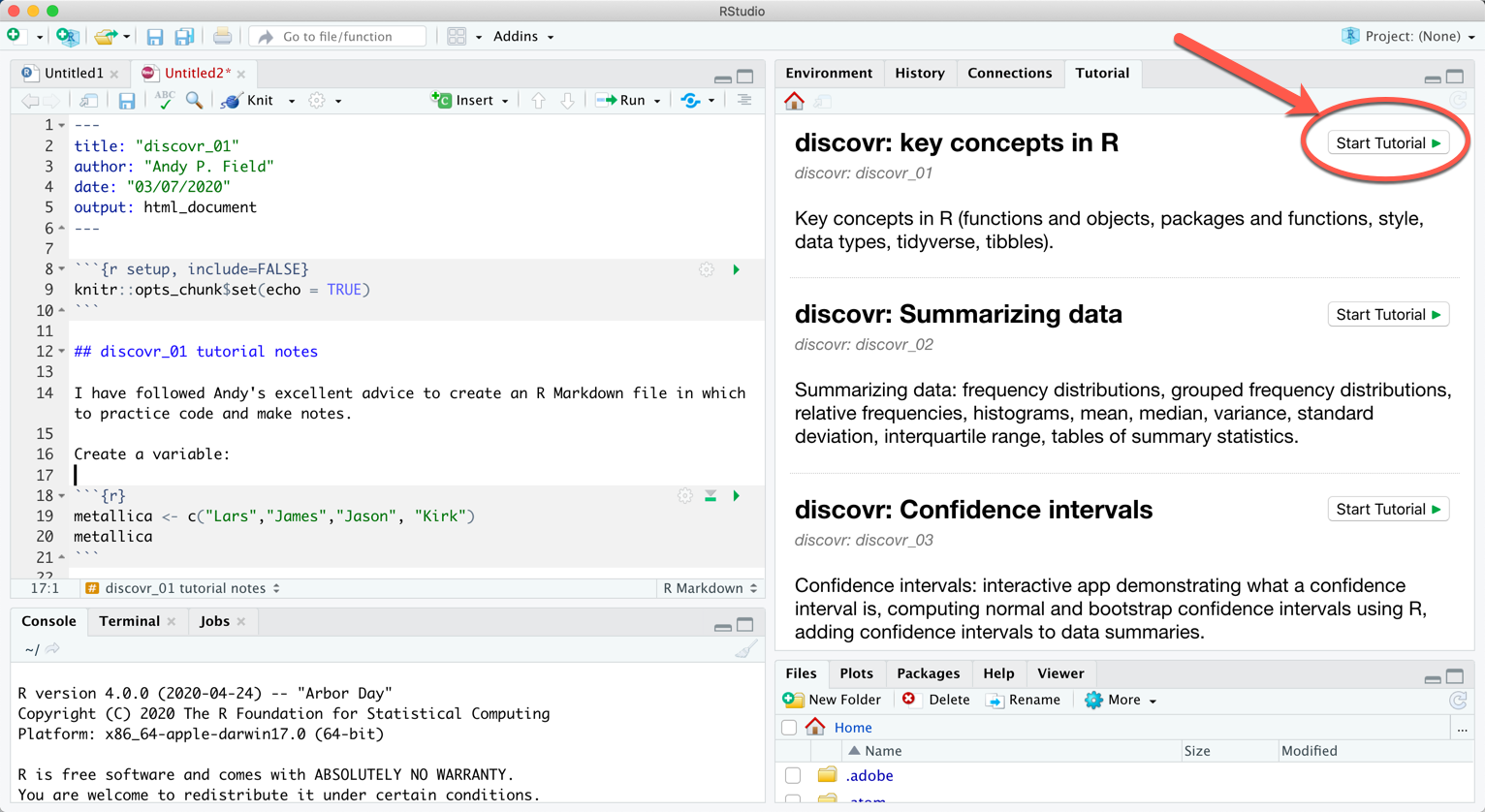
Alternatively, to run a particular tutorial from the console execute:
library(discovr)
learnr::run_tutorial("name_of_tutorial", package = "discovr")
and replace “name of tutorial” with the name of the tutorial you want to run. For example, to run tutorial 3 (for Chapter 3) execute:
learnr::run_tutorial("discovr_03", package = "discovr")
The name of each tutorial is in bold in the list above. Once the command to run the tutorial is executed it will spring to life in the tutorial pane.
Suggested workflow
The tutorials are self-contained (you practice code in code boxes) so you don’t need to use RStudio at the same time. However, to get the most from them I would recommend that you create an RStudio project and within that open (and save) a new RMarkdown file each time to work through a tutorial. Within that Markdown file, replicate parts of the code from the tutorial (in code chunks) and use Markdown to write notes about what you have done, and to reflect on things that you have struggled with, or note useful tips to help you remember things. Basically, write a learning journal. This workflow has the advantage of not just teaching you the code that you need to do certain things, but also provides practice in using RStudio itself.
Here’s a video explaining how I suggest using the tutorials.
Other resources
Statistics
- The tutorials typically follow examples described in detail in Discovering Statistics using R and RStudio. That book covers the theoretical side of the statistical models, and has more depth on conducting and interpreting the models in these tutorials.
- If any of the statistical content doesn’t make sense, you could try my more introductory book An adventure in statistics.
- There are free lectures and screencasts on my YouTube channel.
- There are free statistical resources on my websites www.discoveringstatistics.com and milton-the-cat.rocks.
- R for data science by is an open-access book by the creator of the tidyverse (Hadley Wickham). It covers the tidyverse and data management.
- ModernDive is an open-access textbook on and RStudio.
- RStudio cheat sheets.
- RStudio list of online resources.